Welcome to SpikeInterface’s documentation!
SpikeInterface is a Python module to analyze extracellular electrophysiology data.
With a few lines of code, SpikeInterface enables you to load and pre-process the recording, run several state-of-the-art spike sorters, post-process and curate the output, compute quality metrics, and visualize the results.
If you use SpikeInterface, you are also using code and ideas from many other projects. Our codebase would be tiny without the amazing algorithms and formats that we interface with. See them all, and how to cite them, on our references page. In the past year, we have added support for the following tools:
Bombcell. Bombcell: automated curation and cell classification of spike-sorted electrophysiology data. (docs)
SLAy. SLAy-ing oversplitting errors in high-density electrophysiology spike sorting (docs)
Lupin, Spykingcicus2 and Tridesclous2. Opening the black box: a modular approach to spike sorting (docs)
RT-Sort. RT-Sort: An action potential propagation-based algorithm for real time spike detection and sorting with millisecond latencies (docs)
MEDiCINe. MEDiCINe: Motion Correction for Neural Electrophysiology Recordings (docs)
UnitRefine. UnitRefine: A Community Toolbox for Automated Spike Sorting Curation (docs)
If you would like us to add another tool, or you would like to integrate your project with our package, please open an issue.
Overview of SpikeInterface modules
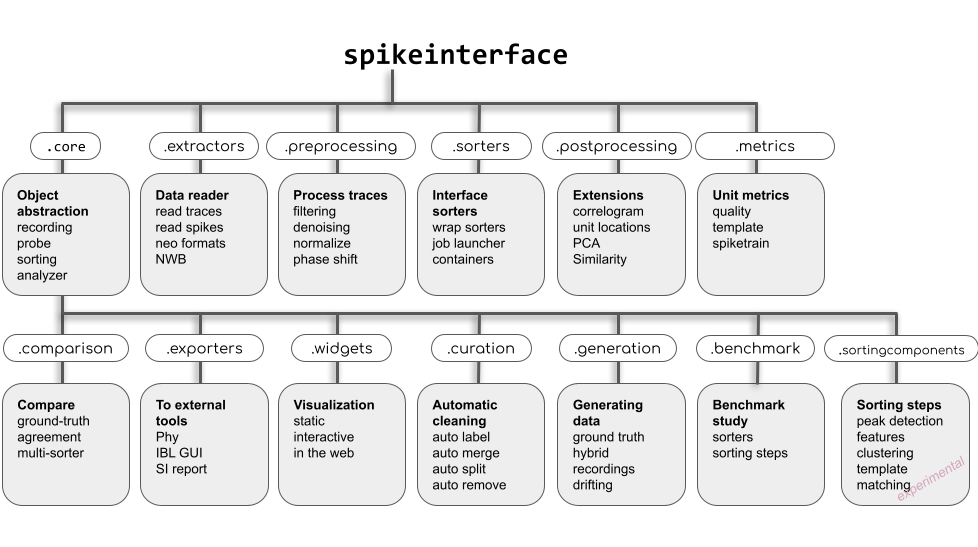
SpikeInterface is made of several modules to deal with different aspects of the analysis pipeline:
read/write many extracellular file formats.
pre-process extracellular recordings.
run many popular, semi-automatic spike sorters (kilosort1-4, mountainsort4-5, spykingcircus, tridesclous, ironclust, herdingspikes, yass, waveclus)
run sorters developed in house (lupin, spykingcicus2, tridesclous2, simple) that compete with kilosort4
run theses polar sorters without installation using containers (Docker/Singularity).
post-process sorted datasets using th SortingAnalyzer
compare and benchmark spike sorting outputs.
compute quality metrics to validate and curate spike sorting outputs.
visualize recordings and spike sorting outputs in several ways (matplotlib, sortingview, jupyter, ephyviewer)
export a report and/or export to phy
curate your sorting with several strategies (ml-based, metrics based, manual, …)
offer a powerful desktop or web viewer in a separate package spikeinterface-gui for manual curation that replace phy.
have powerful sorting components to build your own sorter.
have a full motion/drift correction framework (See Motion/drift correction)
Other resources
To get started with SpikeInterface, you can take a look at the following additional resources:
- spiketutorials is a collection of basic and advancedtutorials. It includes links to videos to dive into the SpikeInterface framework.
- SpikeInterface Reports contains several notebooks to reproduce analysisfigures of SpikeInterface-based papers and to showcase the latest features of SpikeInterface.
- The 2020 eLife paper introduces the concept and motivation andperforms an in-depth comparison of multiple sorters (spoiler alert: they strongly disagree with each other!).Note: the code-base and implementation have changed a lot since the “paper” version published in 2020.For detailed documentation we therefore suggest more recent resources, like this documentation and
spiketutorials.